Key Takeaways
- The paper provides a practical blueprint for an open-source, FAIR-compliant database for sensitive human microbiome data, using the Supabase platform.
- A multi-step procedure involving data anonymization, pseudonymization, and controlled access is crucial to balance the FAIR principles with privacy laws like GDPR.
- Integrating a Large Language Model (LLM), such as ChatGPT, can significantly democratize access to complex scientific data by allowing users to query the database with natural language.
TL;DR
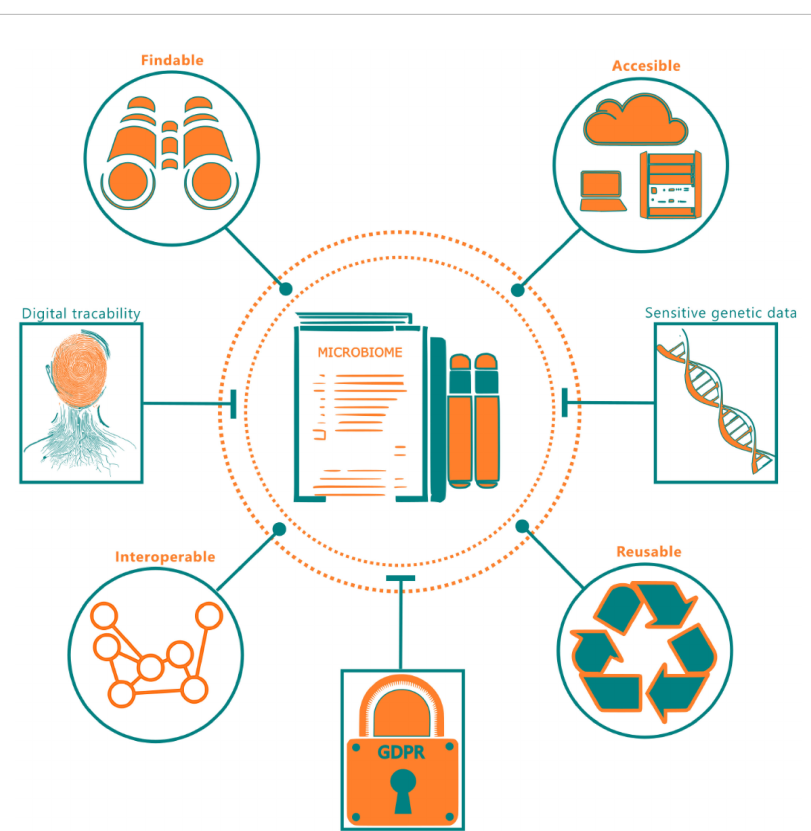
Sharing human microbiome data is crucial for advancing research, but it's a major challenge. The data needs to be Findable, Accessible, Interoperable, and Reusable (FAIR) for collaboration. However, this data is extremely sensitive, containing personal health information and genetic material, which falls under strict privacy regulations like GDPR. This creates a fundamental conflict: how can scientists share data openly while guaranteeing the privacy and security of the human donors? This paper presents a complete open-source solution. The authors developed a FAIR-compliant database using the Supabase platform that implements a robust workflow. Raw data is processed to remove human DNA, and personal information is anonymized. A key innovation is the integration of a Large Language Model (LLM), which acts as a user-friendly interface. This allows researchers and even non-experts to query the complex database using natural language, making the valuable data more accessible than ever before.
Why Does It Matter?
This paper offers a timely and practical solution to a major bottleneck in microbiome research: sharing sensitive data safely and effectively. It provides researchers with a replicable framework for building their own FAIR-compliant, privacy-preserving databases. By demonstrating the use of accessible tools like Supabase and the integration of an LLM for enhanced accessibility, it lowers the barrier to entry for robust data management, fostering greater collaboration and accelerating scientific discovery.
FAIR-GDPR Nexus
This paper explores the critical nexus between FAIR principles and GDPR regulations, revealing a fundamental tension when handling sensitive human microbiome data. It argues that a direct, unfiltered application of the FAIR framework (promoting data reuse via rich metadata) is incompatible with GDPR’s privacy mandates, as it risks patient re-identification. The core insight is that data must be made GDPR-compliant *before* it can be made FAIR. The proposed solution is a GDPR-first processing pipeline that utilizes robust anonymization, pseudonymization, and the removal of host DNA to protect privacy. This ensures the goals of open science are harmonized with the legal and ethical imperative of data protection, creating a practical pathway for compliant and reusable scientific datasets.
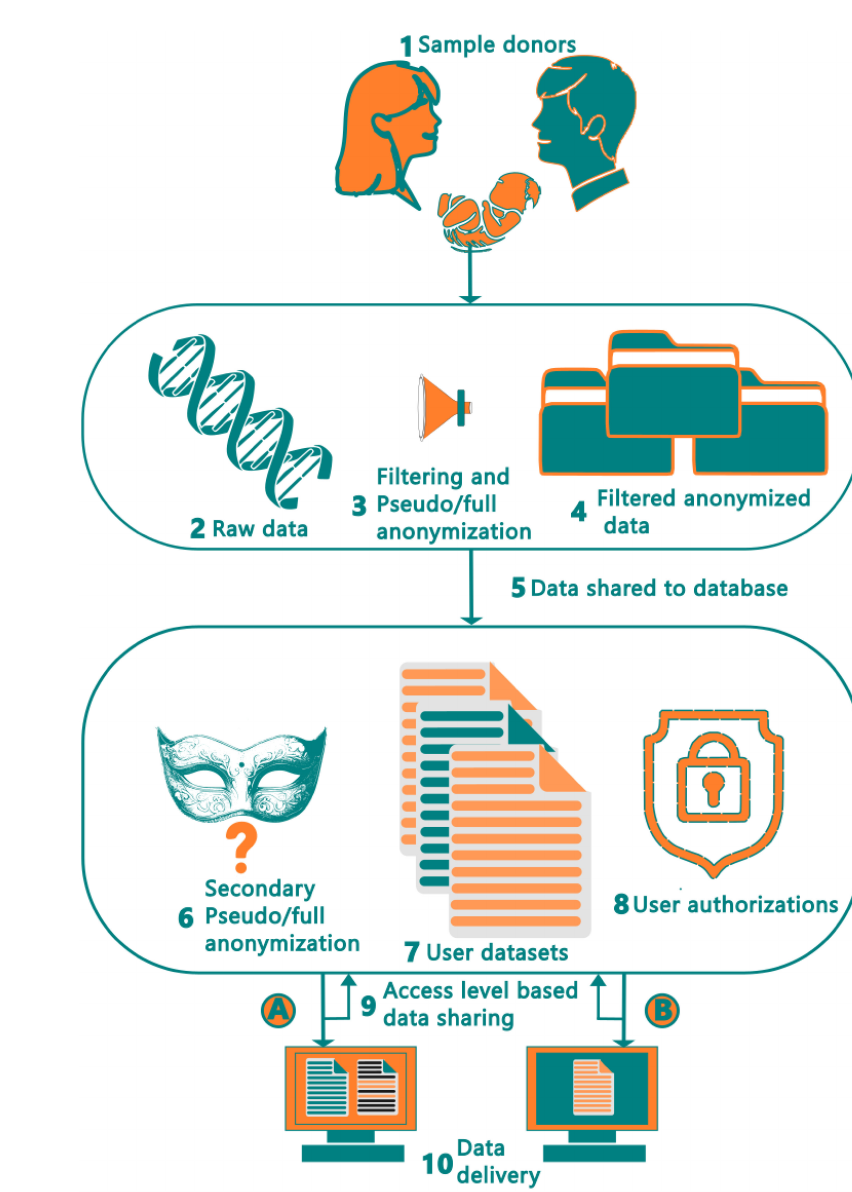
Anonymize Pipe
An 'Anonymize Pipe' conceptualizes a critical, multi-stage data sanitization workflow, essential for reconciling FAIR data sharing principles with stringent privacy regulations like GDPR. This pipeline's initial step involves a technical de-identification of raw metagenomic data by filtering out and removing contaminating human host DNA, which is a direct and sensitive personal identifier. Subsequently, the process would employ robust pseudonymization, replacing direct personal details in the associated metadata with unique, non-reversible subject IDs. This crucial action permits researchers to link different samples from the same individual over time without compromising their identity. The final stage would apply advanced privacy-preserving techniques, ensuring the entire dataset is secure before being made accessible for research.
AI-Access Layer
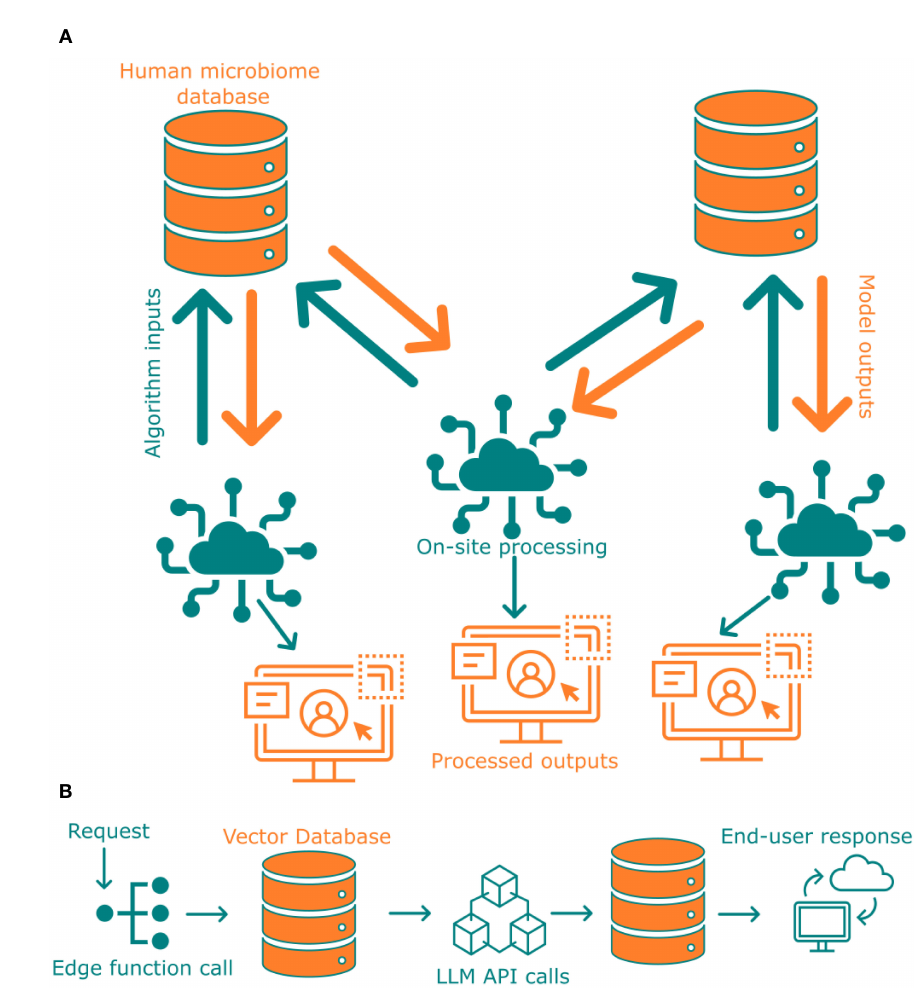
The 'AI-Access Layer' conceptualizes a pivotal framework for democratizing scientific data access. It leverages a Large Language Model (LLM) powered by ChatGPT, integrated through Supabase edge functions and a vector database, to act as an intelligent conversational intermediary. This architecture enables non-expert interaction, allowing users to query the complex, FAIR-compliant microbiome database using natural language instead of specialized code. The layer's primary function is accessibility enrichment, moving beyond technical availability to provide true cognitive and practical access for a broader audience, including students and the public. By translating user questions into actionable database queries, it transforms a static data repository into a dynamic knowledge discovery platform, significantly lowering the barrier to entry for engaging with valuable scientific findings.
Practical Limits
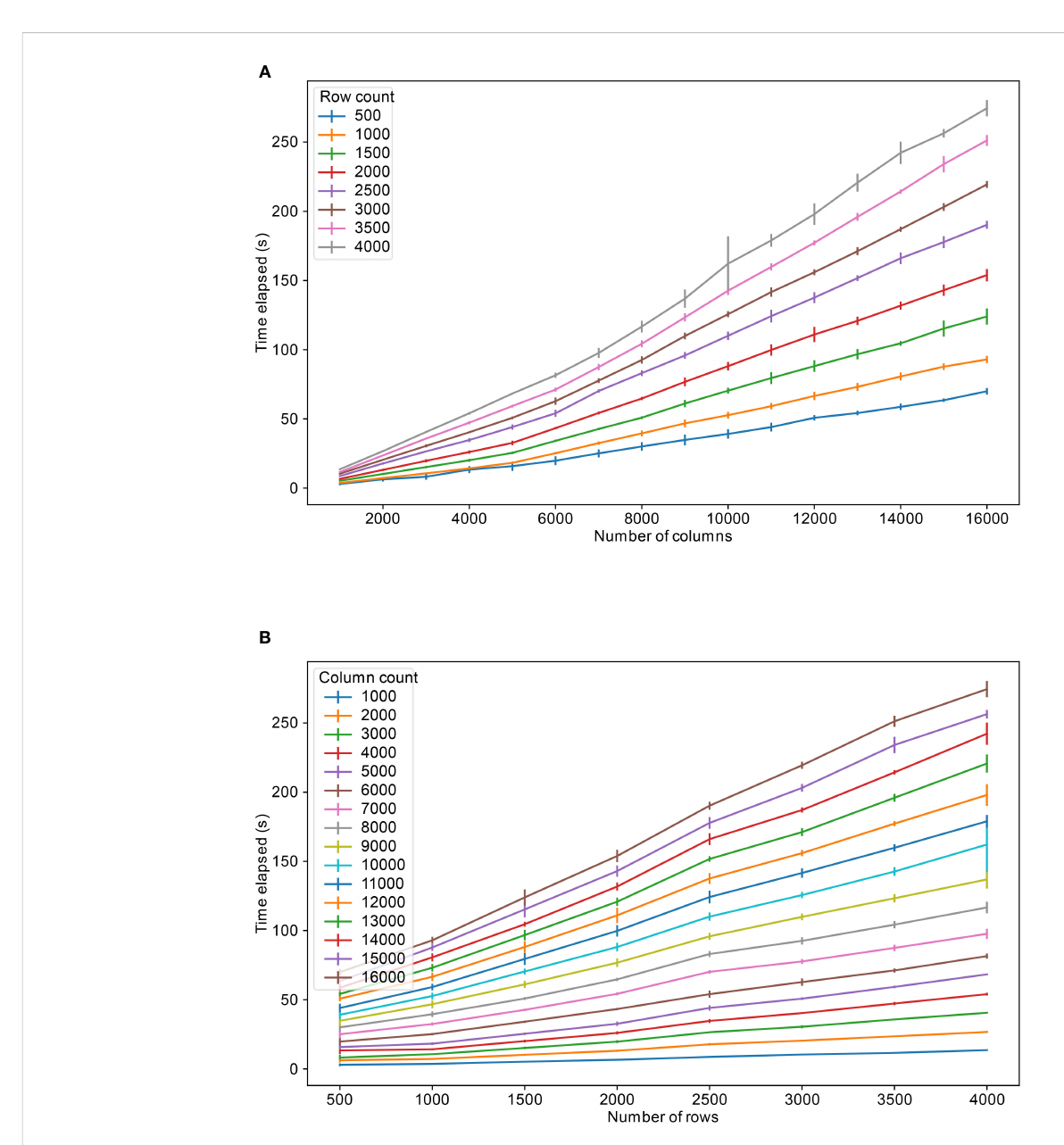
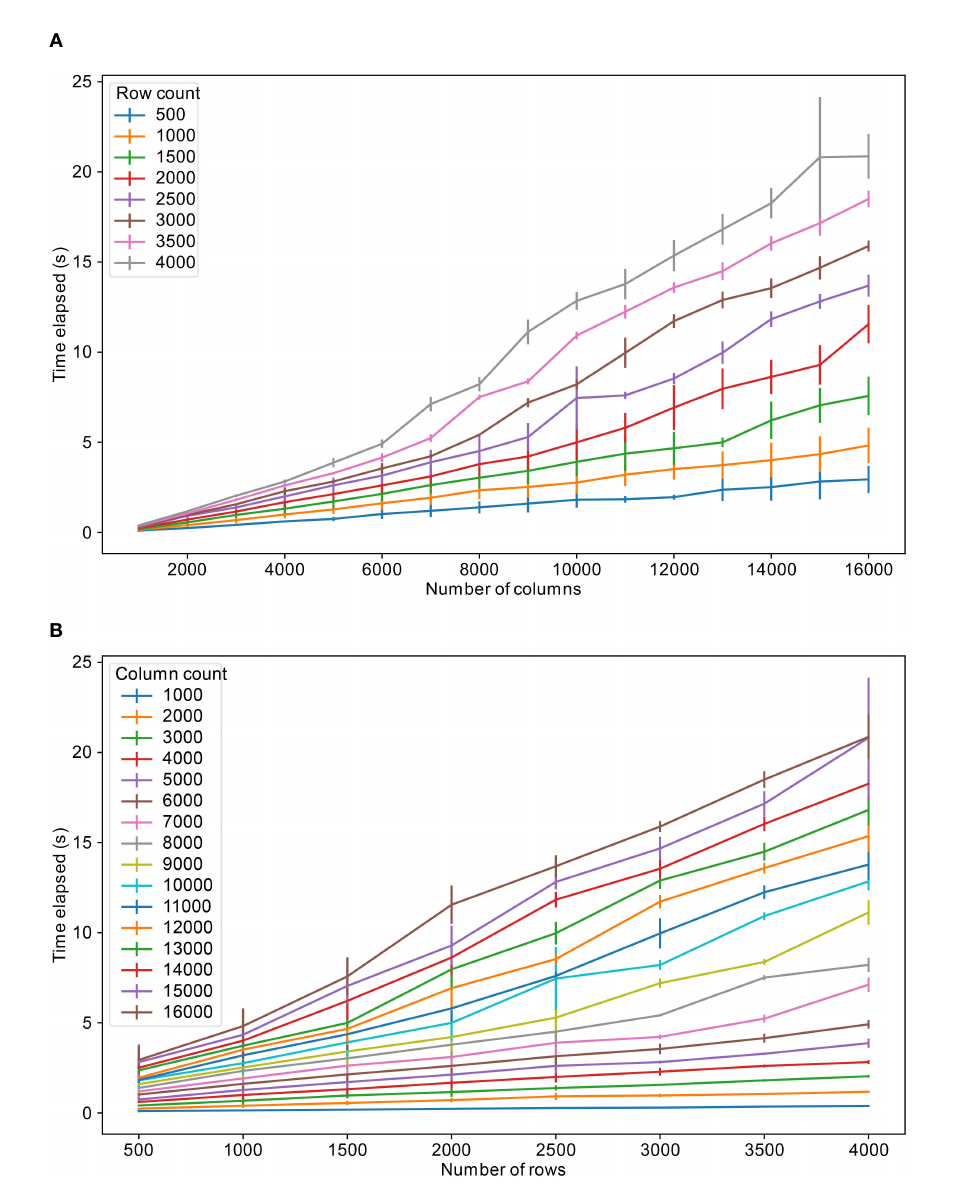
The primary practical limit in developing FAIR-compliant microbiome databases is the inherent tension between data transparency and privacy regulations like GDPR. The presence of sensitive, potentially re-identifiable human DNA within microbe samples makes a strict application of FAIR principles impossible without breaching privacy. This necessitates a trade-off where data utility is deliberately sacrificed for security through imperfect anonymization techniques. Technical limitations, such as database platform constraints and performance bottlenecks with large datasets, further compound the issue. Moreover, the successful implementation and maintenance of such systems demand significant technical expertise and financial investment, creating substantial barriers. Ultimately, the system's design is a compromise, balancing open science ideals against the non-negotiable ethical and legal imperative to protect donor privacy.
Federated Future
The paper envisions a 'Federated Future' for microbiome research, skillfully navigating the conflict between data sharing and privacy. This future does not rely on centralizing sensitive data but on a distributed network of FAIR-compliant, interoperable databases. The crucial insight is to bring computation to the data through federated learning, where algorithms process information locally. Only the anonymized models or analytical results are shared, preserving patient confidentiality under strict regulations like GDPR. This model represents a paradigm shift, enabling global scientific collaboration without a central data repository. The proposed FAIR-compliant database is a critical building block for this secure, collaborative future, harmonizing research advancement with robust privacy protection.
More Figures (4)